V.0 Tutorial 0: Finishing the installation
Once section II is done, the installation is almost finished and you can start the tutorials. Most of section is about configuring the right files path, but inside the NeuroML models, there are also some paths to be configured. The files pointed by the paths to be configured are paramount (see the note below).
Run NMLPlay as explained in section II.B of this user guide. Click on the Environment tab. You should be able to see a panel corresponding to Figure 2.
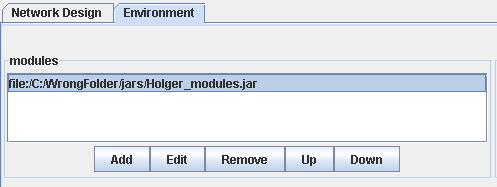
Figure 2: configuring the path of the modules file.
This path might be completely inappropriate. Edit it by clicking on the Edit button at the bottom of the Environment panel. A dialog box will open. Browse through the files tree and select the path of "Holger_modules.jar" .
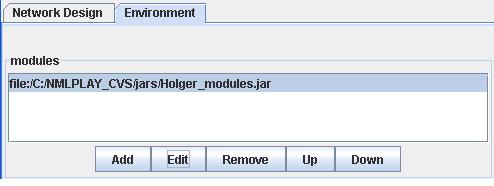
Figure 3: The path of the modules file has been set up.
On my machine, your new root path should appear instead of NMLPLAY_CVS/jars, which is the new root path on my machine.
You should then save the file as BasicModel.xml in your folder RootPath/jars/BasicModel.xml. In order to do so click on File in the menu and then Save. Select the right path in the newly opened dialog box (Figure 4).
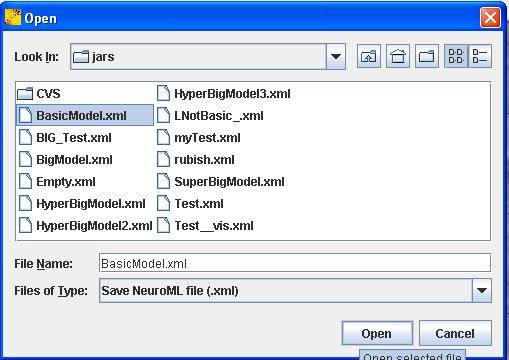
Figure 4: The save dialog box.
You will then need to configure all your other NeuroML file. In order to open a NeuroML file, click on File in the menu, and then open. Select the path of the next NeuroML file. Do change the path of the module file appropriately, save the neuroML file as explained above. Do proceed will all the remaining NeuroML files in the likewise.
Note: The module files contain a set of algorithms representing some biological and physical mechanisms. In particular, the module file sample "Holger_modules.jar" contains:
- Several neuron models. An integrative model, Hodgkin-Huxley models with or without synapses.
- Learning synapses, whose weight depends on the frequency of the spiking activity.
- Different filters for the projections, which help to determine how neurons should be interconnected. These filtering modules are referred as a "destMethod" or a "sourceMethod". A "sourceMethod" basically filters the neurons in the source population which request a connection. A "destMethod" filters the requests received by the destination population from the source population.
- A set of devices to monitor the simulation, such as volt/ampere/multi-meters, a device to monitor the spiking activity of the whole network. These devices are implemented in some modules called "moduleAttachment".
- A rendering environment which determines how the results of the attachments are displayed. The rendering modules are called "moduleProbe".
Later the user construction of modules is explained in tutorial 8.
Even if modules define some mechanisms, most of the time, they allow the user to input some parameters which specify more precisely their behaviour. Therefore you need to instantiate the modules, and define these parameters.