Introduction
NeuroGems is a collection of neuroinformatics software modules for data and modelling in neuroscience. The emphasis is on tools for collaboration, visualisation and use of open XML standards.
The project was an escience pilot project funded by the UK MRC and BBSRC and based at the Universities of Edinburgh, Newcastle and UCL running from 2002 - 2005.
Follow-on projects are being developed by Textensor including:
|
Textensor Limited is a software company started by former Edinburgh University researchers which specialises in creating web tools for collaborating on documents and also undertakes a range of software consultancy in modeling and simulation of biological systems. In April 2008 Textensor launched the A.nnotate service. The PublicationsList.org service, launched April 2007 is now used by over 5000 researchers world wide to improve visibility of their research. |
||||||||
|
Axiope
|
|||||||||
|
The Axiope project has developed software for cataloging and publishing scientific data, a user friendly route to XML and federated databases. The project resulted in a spin-out company which has developed a commercial version of the software. (Founded by Fred Howell and Robert Cannon, University of Edinburgh, with commercial exploitation by Nigel Goddard, Axiope Ltd). |
||||||||
|
NeuroML
|
|||||||||
|
The NeuroML project is a collaboration which includes support for defining new XML languages for models and experimental data in neuroscience. (Fred Howell, University of Edinburgh). |
||||||||
|
neuroConstruct
|
|||||||||
|
The neuroConstruct application provides an interface for creating 3D models of networks of biologically realistic neurons, which can be mapped onto the NEURON or GENESIS environments for simulation. (Padraig Gleeson, Angus Silver, Volker Steuber, UCL). |
||||||||
|
NClamp
|
|||||||||
|
NClamp is a suite of tools for data acquisition software for electrophysiology. (Jason Rothman, A. Silver Lab, UCL). |
||||||||
|
RatBrain
|
|||||||||
|
The RatBrain project has developed a database and tools for integrating 3D neural reconstructions from different areas of the rat brain performed using the Neurolucida and Accustage systems. (collaboration between Edinburgh University and Rutgers University). |
||||||||
|
3D Atlas tools for Macaque
|
|||||||||
|
The 3D atlas project, in collaboration with Rolf Kötter, Duesseldorf is developing software for visualising connectivity and brain atlas data from the CoCoMac database of Macaque cortex. (collaboration between Edinburgh University and the Kötter lab). |
||||||||
|
Catacomb
|
|||||||||
|
Catacomb is a collection of software for investigating multi level models of ion channels, compartmental neurons and networks of spiking cells. (Dr Robert Cannon, Edinburgh). (Development was supported by this escience project in 2002-3. |
||||||||
|
Abstracted Protein Simulator
|
|||||||||
|
The abstracted protein simulator is an experimental meso-scale monte-carlo simulator for modelling formation of multi-protein complexes. (Dr Fred Howell and Eilidh Grant, Edinburgh University) |
||||||||
|
Parallel discrete event simulation
|
|||||||||
|
The neosim project includes a parallel discrete event simulation kernel for running models of spiking neurons on a cluster of workstations. Models are specified using NeuroML, and visualised using Java2D. |
||||||||
|
Growing artificial neural connectivity
|
|||||||||
|
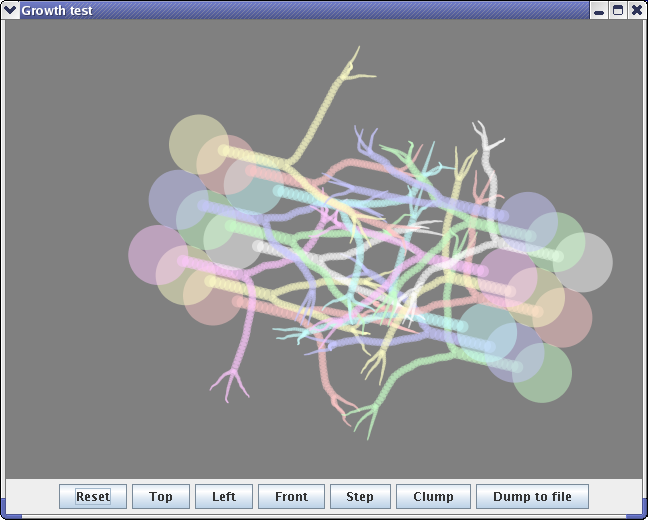
The neural growth software grows neurons into a volume of simulated tissue to generate packed volumes of neural processes. The aim is to generate known test data for automated routines which deduce connectivity from volumetric EM slices. |
||||||||
|
Automatically segmenting EM images
|
|||||||||
|
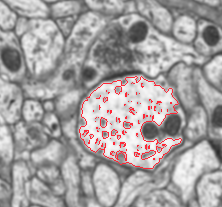
The voxel (3D) and marching (2D) packages use fast marching / level set techniques to extract outlines of neural processes from electron microscope slices of brain tissue. The long term goal is to further develop techniques for automatically obtaining quantities of high resolution connectivity data from volumetric EM. The quantities of image data involved can be enormous for even small volumes of brain, making distributed analysis across large grids of computers essential once the algorithms reach a quality threshold. |
||||||||
|
Utility: A generic Python XML-Object serializer
|
|||||||||
|
The nmlparser utility is a library for simulator developers using Python who wish to serialise their complex object models into elegant XML. It uses Python reflection in combination with a simplified XML schema format to minimise the amount of developer effort to generate or parse XML formats (including MorphML) while ensuring that the XML produced is more elegant, concise and language- independent than existing object serialisers. |
||||||||
|
Utility: Spidering PLOS Biology for text mining applications
|
|||||||||
|
The fetchplos utility contains a script for downloading the full text of all papers in XML from the PLoS Biology journal website. It is useful to have large numbers of papers available locally as a starting point for text and data mining experiments, and the script could be easily adapted to download local copies of papers from the other PLoS journals, as well as papers on PubMed. |
||||||||